Welcome to VariantExplorer!
The goal of VariantExplorer is to facilitate identification of clinical significance interpretation discrepancies in ClinVar (http://www.ncbi.nlm.nih.gov/clinvar/), a submitter-driven repository that archives reports of the relationships among genomic variants and phenotypes submitted by clinical laboratories, researchers, clinicians, expert panels, practice guidelines, and other groups or organizations. Given the large number of submitters to ClinVar, many variants have interpretations from multiple submitters and those interpretations may not always agree.
By displaying how the full set of variant interpretations from a specific submitter compares to all other submitters (or to another specific submitter), VariantExplorer helps users view the types and levels of discrepancies in ClinVar. The submitter-specific Clinical Significance Breakdown Tables (seen below) displays pair-wise counts of discrepant interpretations, including confidence discrepancies (such as Benign vs Likely benign or Pathogenic vs Likely pathogenic). For example, the table below indicates there are 12 variants in ClinVar interpreted as Likely benign by Submitter A and interpreted as Uncertain significance by Submitter B. By displaying the discrepancies in this manner, VariantExplorer hopes to facilitate resolution of interpretation discrepancies.
Clinical Significance Breakdown Table example
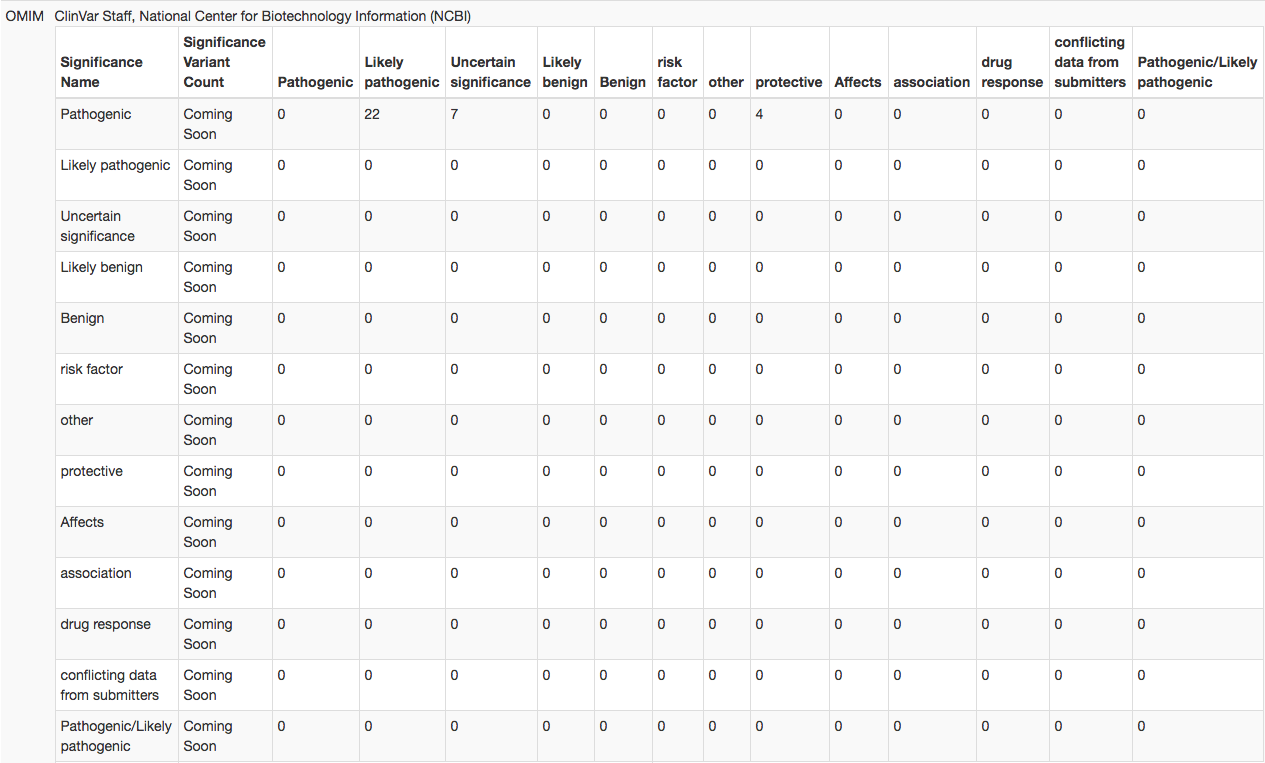
The discrepancy data in VariantExplorer can be viewed from four different approaches:
Search By Submitter
This option allows users to view all discrepancies with regard to a specific ClinVar submitter. Selecting a ClinVar submitter navigates to a Submitter by Submitter Summary table of all submitters with interpretations that are discrepant with the submitter of interest. The discrepancy counts are broken into Confidence Discrepancy and Conflict. Below the summary table are the Clinical Significance Breakdown Tables of each submitter-submitter pair listed in the Submitter by Submitter Summary table. Clicking the counts in any Clinical Significance Breakdown Table displays the variants with clinical significance discrepancies and summary information about each submission, such as asserted condition and date last evaluated. Selecting the variant name will direct a user to the variant page in ClinVar.Show Submitter Mega Table
This option allows users to view discrepancy counts between all submitters in ClinVar. Each submitter is assigned a lab number and hovering over a lab number or discrepancy count displays the full name of that ClinVar submitter. Clicking the discrepancy counts in any cell will display those variants of interest below the table.Search By Significance
A Clinical Significance Breakdown Table of all discrepancies in ClinVarSearch By Variant
A dropdown list of all variants in ClinVar with clinical significance discrepancies